软硬兼施 | Leica打造的3D肿瘤球培养的解决方案
 生命诞生于3d环境,所以传统的2d细胞培养方法,虽然可以保证细胞的生长和对外界刺激产生生理反应,但是和实际生活环境的巨大差异会导致大部分生理功能受限。比如,2d培养环境下的细胞间的相互作用是xy轴的,只是细胞层之间的作用,缺乏z轴方向的影响,这样就没有养料、氧气和外界刺激物(如:药物处理)的梯度渗透作用。而3d培养环境下细胞可以变成细胞球,从而可以体现出z轴细胞之间的力作用和各种物质的渗透梯度,和体内的差异相较2d细胞层会大大缩小,从而提高体外细胞实验的准确性。
生命诞生于3d环境,所以传统的2d细胞培养方法,虽然可以保证细胞的生长和对外界刺激产生生理反应,但是和实际生活环境的巨大差异会导致大部分生理功能受限。比如,2d培养环境下的细胞间的相互作用是xy轴的,只是细胞层之间的作用,缺乏z轴方向的影响,这样就没有养料、氧气和外界刺激物(如:药物处理)的梯度渗透作用。而3d培养环境下细胞可以变成细胞球,从而可以体现出z轴细胞之间的力作用和各种物质的渗透梯度,和体内的差异相较2d细胞层会大大缩小,从而提高体外细胞实验的准确性。最近的统计表明每年会有370万新增癌症病例在欧洲发生,这些疾病会导致20%的死亡率。利用体外培养的肿瘤细胞来高通量的筛选新的有效的药物对于挽救癌症病人非常重要,而3d培养的肿瘤细胞球是合适的模型,但是这种细胞模型也有其瓶颈。因为3d细胞球的培养方案不统一,而且生长环境较之2d复杂,所以最终养成的球体异质性会较之2d细胞层严重,而药物的筛选需要有尽量均一的样本,这样最终得到的药物敏感数据才准确可信。为了解决这样的情况,需要对培养出来的肿瘤球进行挑选,然后放到孔板中,但是传统的挑选方法是用人为的用肉眼挑选,这样会导致效率的低下,而且个人的差异会导致标准不统一,再者如果直径500微米一下的球体挑选难度就会更大。此外,对3d球体在镜下和摄像头下的成像也容易受到光源、培养基浓度和培养器皿的形状而影响,从而更加增加观察者肉眼判读的不统一性。1、2、3、4、5、6、7、8、9、10
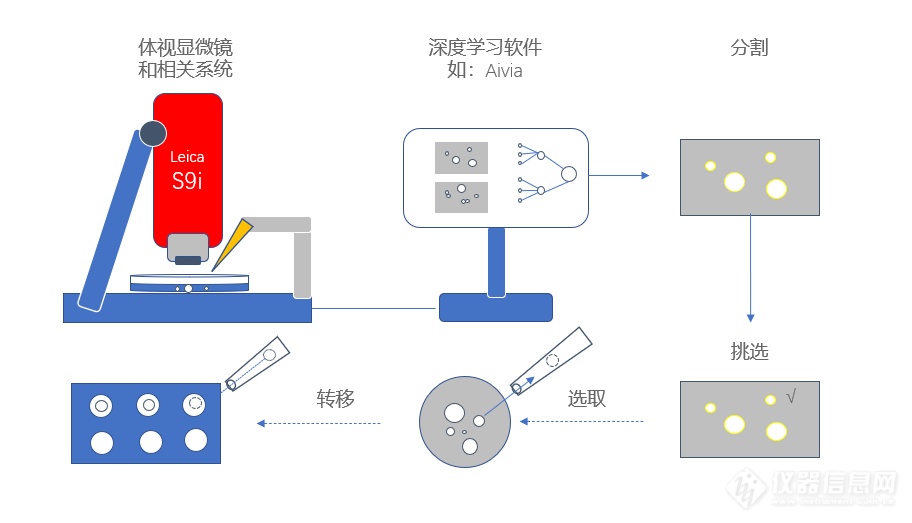
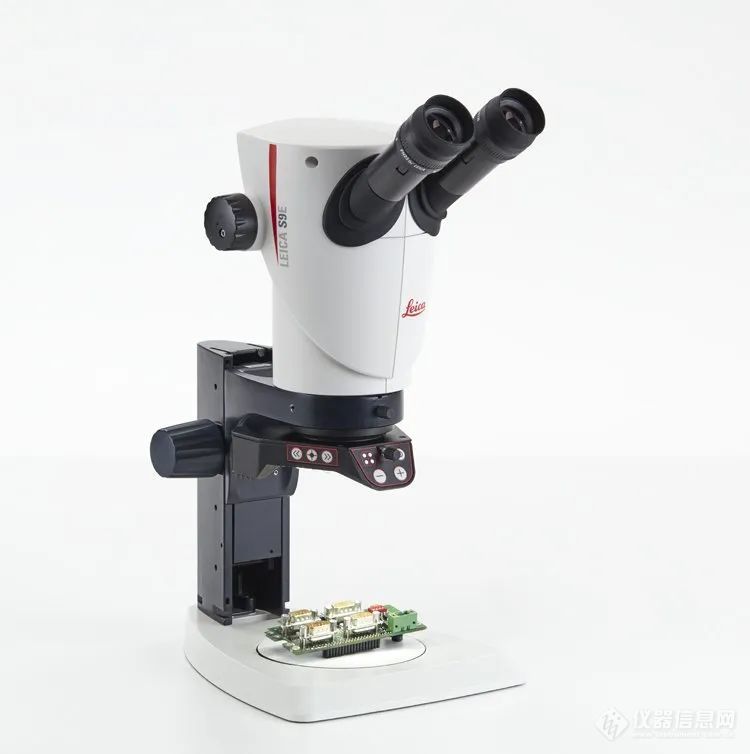
 leica s9i体视镜独有的fusionoptics融合光学系统,可以同时兼顾长景深和高分辨率,而3d球体因为有z轴的深度,所以使用这种技术可以保证完整而清晰的观察到球体。且放大倍率可以到55x,实现9:1的变倍比,实现总览到细节的快速切换。此外,该机型搭配了10mp的摄像头。光学素质和高成像的分辨率可以方便为深度学习软件来分析。再加之有122 mm工作距离,从而有足够的空间来搭配上图中的自动化显微操作系统和注射器。
leica s9i体视镜独有的fusionoptics融合光学系统,可以同时兼顾长景深和高分辨率,而3d球体因为有z轴的深度,所以使用这种技术可以保证完整而清晰的观察到球体。且放大倍率可以到55x,实现9:1的变倍比,实现总览到细节的快速切换。此外,该机型搭配了10mp的摄像头。光学素质和高成像的分辨率可以方便为深度学习软件来分析。再加之有122 mm工作距离,从而有足够的空间来搭配上图中的自动化显微操作系统和注射器。
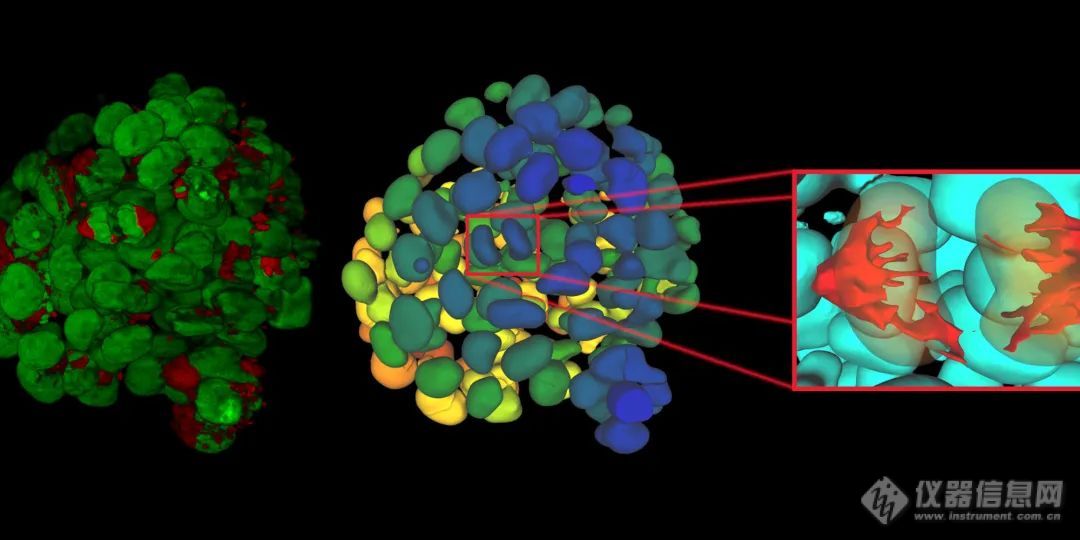
细胞球的精确分割是整个细胞选择培养流程的重中之重。在自动选择过程中需要兼顾灵敏性及准确度问题,以排除尺寸过小、过大或外形不规则的细胞球个体。由于细胞球通常不会彼此接触或重叠——这一特点恰巧符合u-net 神经网络的设计,因此通过u-net 神经网络训练的深度学习模型可以很好的解决自动地选择细胞球这一难题。而同为leica旗下的aivia人工智能图像分析软件,是一款具有处理深度学习模型的软件,此外其还为使用者提供了python 接口,用户可以调用公开的三方模型或分析工具,如发布在github 上的资源,也可以根据实验需要编写符合特定需求的插件,制定一体化的分析流程,提高整体分析流程的效率。
经过这套系统转移后的球体状态如何呢?作者通过leica sp8共聚焦检测了形态和活力相关的标记物(用calcein am、eth-d1和hoechst3342),通过短时间(转移前后马上对比)和长时间(0、24、48小时),发现通过spheroidpicker的转移操作几乎不会影响细胞球的活力。
而这套系统的最终效果如何呢?要求专家和spheroidpicker挑选面积为21,000-29,000 μm2、最小圆度为0.815的小球。对比最终结果,发现spheroidpicker对挑选面积和圆度的范围控制,其操作28次,其中26个小球被成功挑出,25个转移成功,成功率更高。
3d球体是现在最有前途的体外筛药模型,让细胞xyz空间展现出较之2d单细胞层更多更准确的生理功能,但是其培养方法的不统一造成较难获得均一化个体,需要后期的挑选才可以作为可信的筛药对象,而借助能拥有光学素质能兼顾清晰和3d景深leica s9i体视镜加上深度学习软件(leica aivia可以用来开发此功能),可以自动化的来做挑选和转移的工作,从而让3d球体成为可信高效的体外筛药模型。
参考文献:
更多![]()
美谷分子微生物筛选系统 QPix亮相央视晚间新闻 ,助力创新中国
厂商
2024.07.11
推动生物合成学的创新解决方案
厂商
2024.07.11
如何科学延长超速离心机的使用寿命?
厂商
2024.07.08
高效自动化胶囊装填系统:专为临床试验量身定制的优化方案
厂商
2024.07.04